Project Overview
MolecuLens is an educational drug discovery platform I built to help students explore how machine learning can support early stage molecular screening. The platform gives users a guided workflow where they can select a disease target, enter or choose a molecule, and review model predictions in a way that feels clear instead of overwhelming.
I developed this project for chemistry and biology learning at Thompson Rivers University. The main goal was to make advanced drug discovery ideas more accessible in a classroom setting by turning a complex scientific process into a fast and understandable web experience.
Access Project
Use the live project link below to view the public MolecuLens site and project information.
Open MolecuLensFast for learning
Students can move from target selection to model output in a few seconds, which makes the tool practical for lectures, labs, and individual exploration.
Rich scientific output
The platform combines activity prediction, molecular property checks, toxicity screening, and structure visualization in one interface.
Built for education
The interface is designed to explain the workflow visually, so users can connect concepts from class to an actual applied system.
The Learning Problem
Drug discovery is usually taught through theory, diagrams, and limited demonstrations. In practice, the real tools are often expensive, difficult to configure, or not intended for students who are still learning the basics. That creates a gap between understanding the idea of molecular screening and actually seeing how it works.
High technical barrier
Many chemistry students do not work with Python environments, notebooks, or machine learning libraries on a regular basis. That makes it hard to use research tools without heavy setup.
Limited classroom time
Instructors need tools that respond quickly and fit into a teaching session. Slow workflows or unstable toolchains make live exploration difficult.
Weak concept retention
Students understand ideas better when they can test inputs, compare outcomes, and see why one molecule behaves differently from another.
Guided Analysis Workflow
MolecuLens was designed as a guided workflow rather than a loose collection of tools. The interface walks the user through target selection, molecule selection, prediction, and interpretation in a sequence that feels simple and intentional.
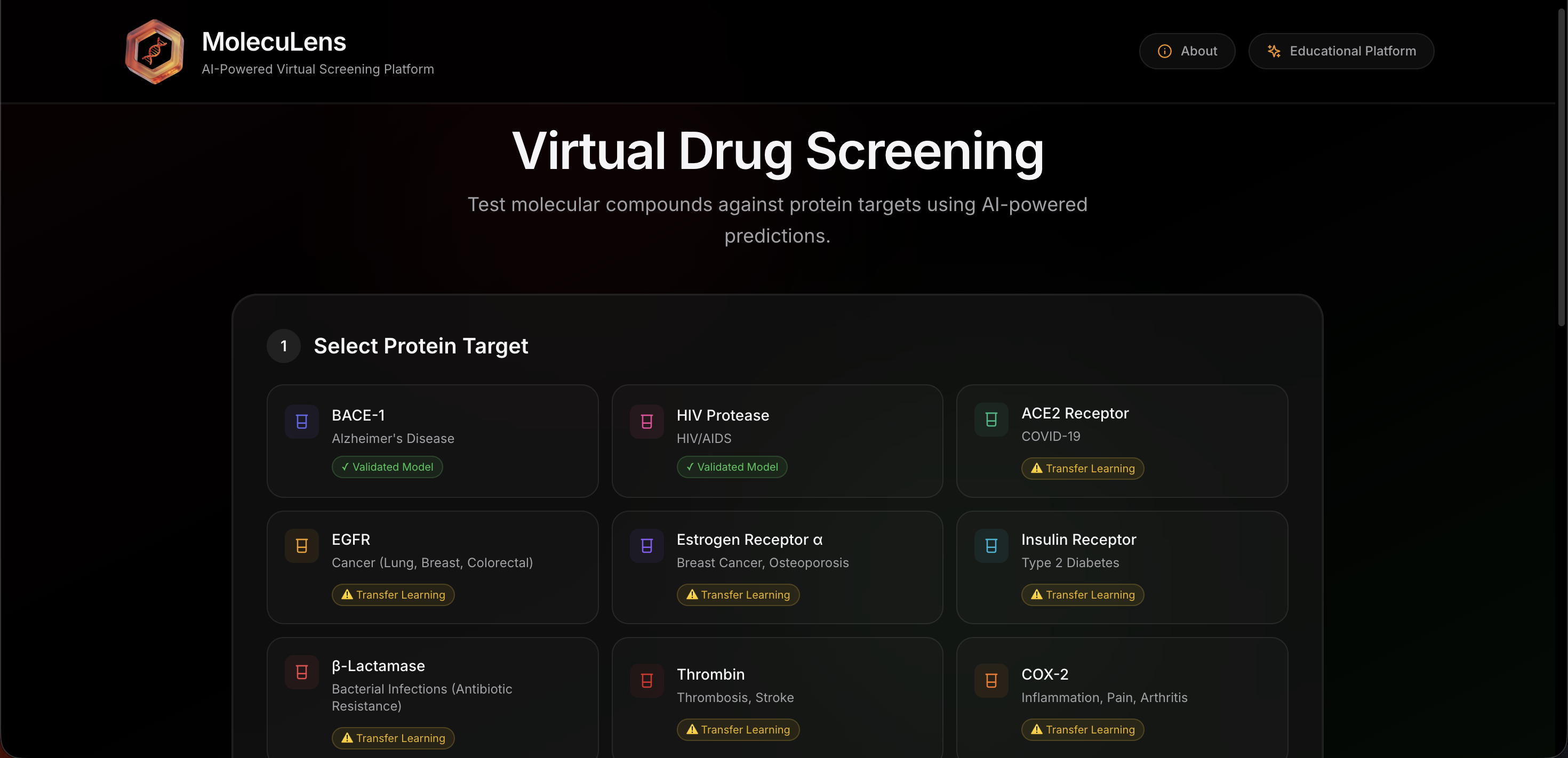
How a user moves through the platform
- Choose a protein target linked to a disease use case.
- Select a sample molecule or enter a custom SMILES string.
- Run the prediction pipeline and wait for the model outputs.
- Review activity, property, toxicity, and structure results.
- Compare molecules side by side when needed.
This flow keeps the focus on interpretation rather than setup. It makes the project useful for teaching because students can spend more time understanding what the results mean.
Supported exploration
The platform supports multiple disease targets and includes example molecules so students can start immediately. This lowers the entry barrier for first-time users while still allowing more advanced testing through custom molecular input.
By combining guided defaults with open input, MolecuLens works well for both structured class activities and open-ended experimentation.
Feature Deep Dive
The value of MolecuLens comes from how its features work together. It is not just a prediction form. It is a compact learning environment that helps users understand the tradeoffs behind a candidate molecule.
Target and molecule selection
The first part of the experience is designed to be fast and clear. Users can choose from supported disease-related protein targets and then work with either preloaded molecules or custom SMILES input. This gives beginners a safe place to start while still supporting deeper exploration.
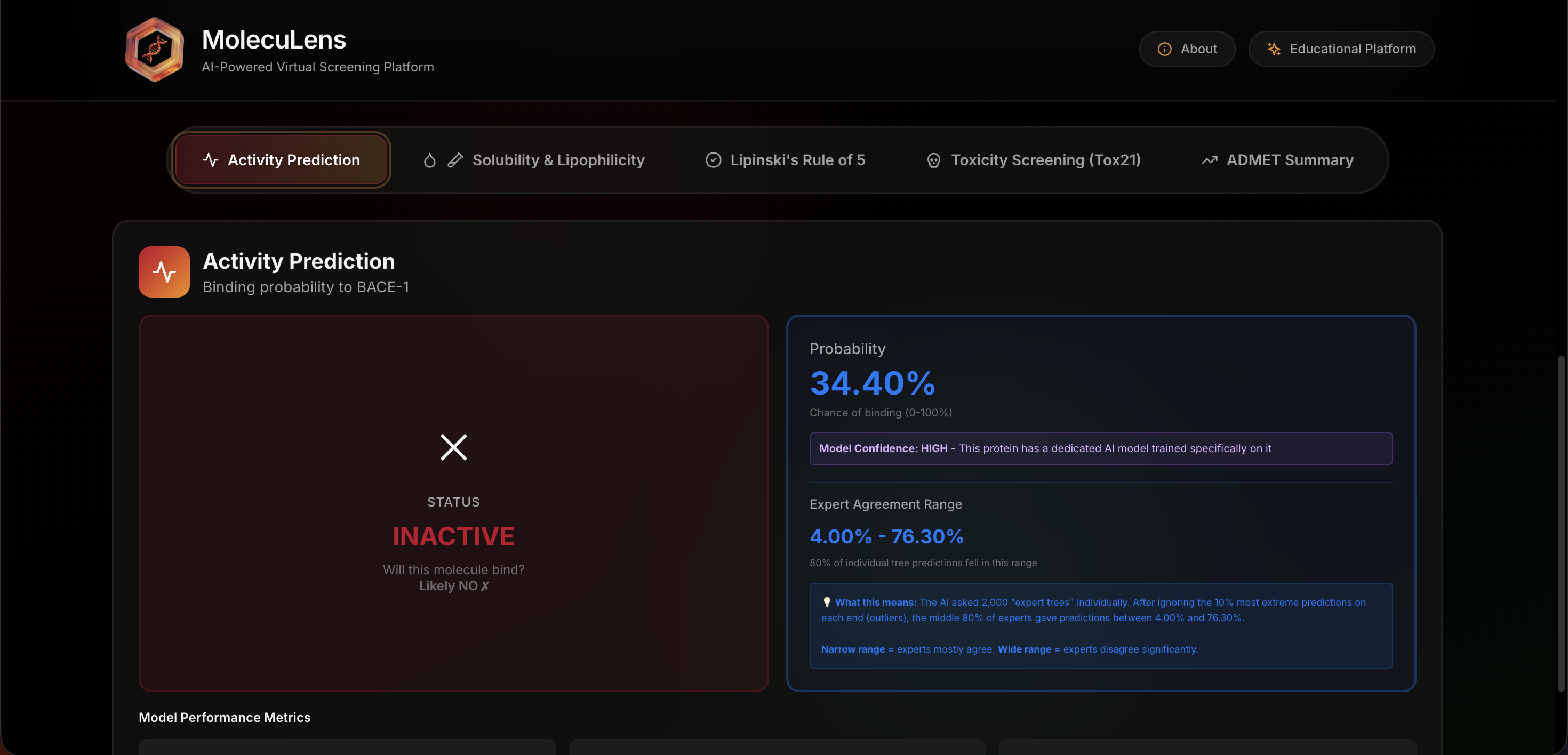
Prediction dashboard
After analysis, the dashboard organizes the output into readable sections. Instead of showing a confusing wall of numbers, it presents activity likelihood, molecular properties, and interpretation cues in a format that is easier to review during class discussion.
Property and safety checks
MolecuLens does more than classify activity. It also reviews solubility, lipophilicity, drug-likeness, toxicity, and related indicators so users can see that a promising molecule is not judged by a single score alone.
2D structure visualization
Structure rendering helps connect abstract molecular notation to a more visual interpretation. This is especially useful for students who are still learning how structural form relates to behavior and predicted performance.
Why the dashboard matters
A major design goal was to keep the results readable. Each prediction type has a clear place in the interface so that users can move from one signal to the next without losing context.
That makes the project useful in both teaching and demonstration settings because it supports quick explanation. An instructor can move through the output logically and explain why certain molecules look more promising than others.
Classroom Value
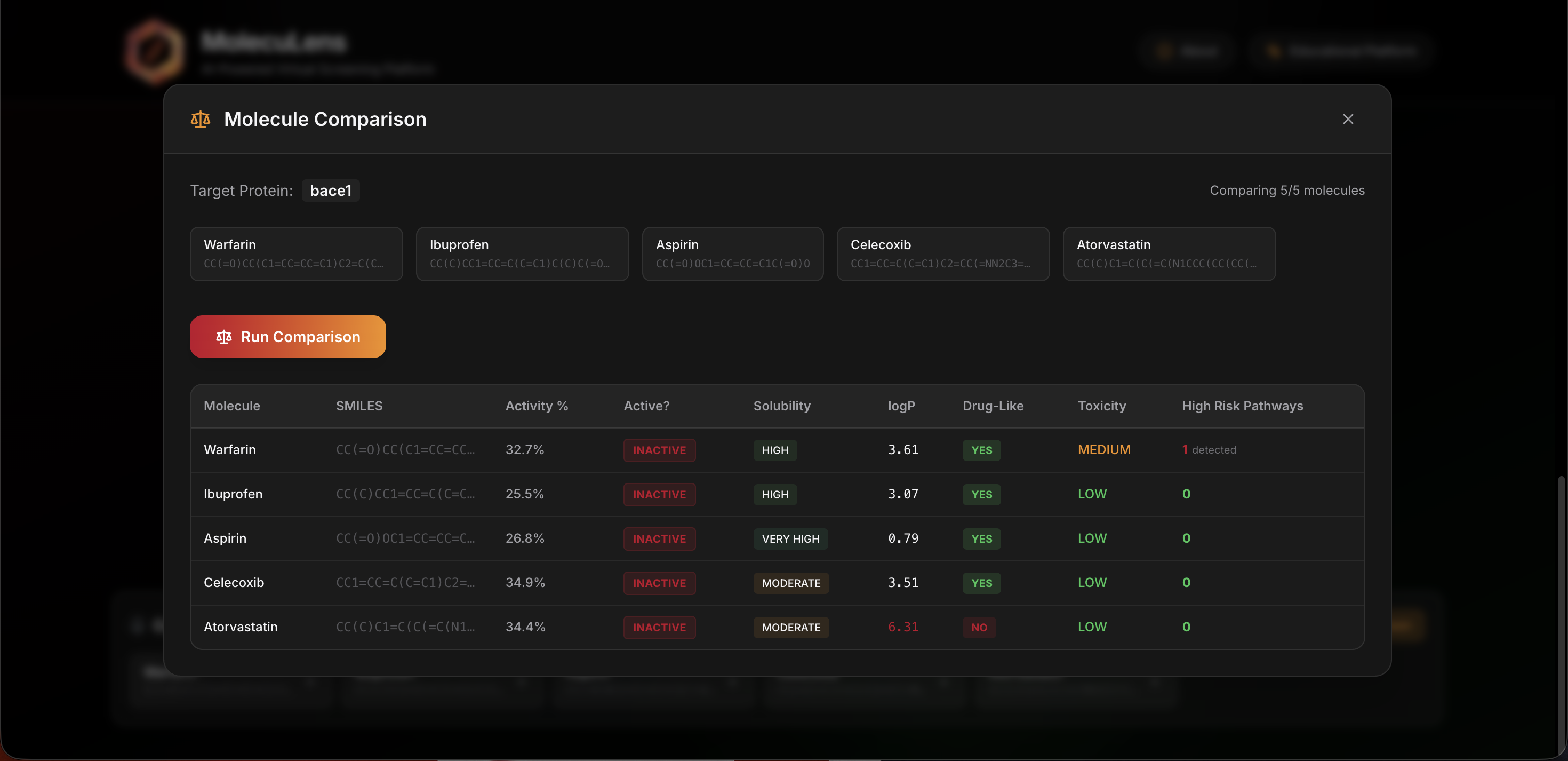
Built for comparison and discussion
One of the most useful features in a classroom setting is the ability to compare molecules side by side. Instead of reviewing one result in isolation, students can look at multiple candidates together and talk through the tradeoffs.
This comparison view helps answer better questions. For example, a molecule might show stronger predicted activity but weaker safety or property scores. Seeing that tension directly makes the lesson more memorable.
Why this helps teaching
- It turns abstract model output into direct comparison.
- It supports assignments that ask students to justify a choice.
- It encourages interpretation instead of passive reading.
Technical Architecture
MolecuLens combines a modern frontend with a Python-based machine learning backend. The architecture was built to keep the user experience smooth while still supporting model inference and chemistry-focused processing.
Frontend layer
Next.js, React, and TypeScript drive the interface. The frontend is responsible for input collection, result presentation, and structure rendering support through RDKit-based visualization.
Backend layer
FastAPI handles the prediction requests and coordinates the analysis pipeline. This keeps the model logic separate from the user interface and makes the system easier to maintain.
ML and chemistry layer
DeepChem, RDKit, scikit-learn, and related chemistry datasets support the activity, property, and toxicity models that power the platform.
Design focus
The technical work was important, but the real challenge was turning the system into something students could actually use without friction. That required careful attention to response time, readability, and how much information the interface should reveal at each step.
The result is a project that demonstrates machine learning in a real use case while still respecting the needs of a teaching environment.
Screenshots

Landing and input flow
This screen introduces the target selection and molecule input flow.

Results dashboard
This view brings together prediction output and interpretation.

Comparison mode
This screen helps users compare multiple molecules side by side.
Results and Learnings
MolecuLens helped me practice a complete project flow that combined user experience design, API architecture, chemistry-focused machine learning, and educational product thinking. It also reinforced a simple lesson: advanced tools only become useful in teaching when they are fast, clear, and easy to approach.
Project outcome
The final platform gives students a practical way to explore drug discovery ideas without needing a full research environment. It makes machine learning feel concrete and usable instead of distant.
What I learned
I learned how to package scientific and machine learning workflows in a way that respects real users. Building for education requires more than model output. It requires structure, clarity, and thoughtful presentation.